Introduction to the archaeacentre package
Richard Stöckl
Archaea Centre Regensburgrichard.stoeckl@ur.de
12 February 2025
Source:vignettes/archaeacentre.Rmd
archaeacentre.RmdBasics
Install archaeacentre
R is an open-source statistical environment which can be
easily modified to enhance its functionality via packages. biocthis
is a R package available via the Github
repository.
Get the latest stable R release from CRAN. Then install the development
of archaeacentre from GitHub
with:
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install(c("remotes", "richardstoeckl/archaeacentre"))Presets for growth curves of microbial growth data
Plot a basic growth curve
# 1. load the package
library(archaeacentre)
# 2. load some test data included with the package
testData <- archaeacentre::growthData
# 3. plot the growth curves in their most basic way, with the presets for "robert":
archaeacentre::plotGrowthCurve(testData, timepoint, concentration, grouping = c("timepoint", "organism"), organism, type = "robert")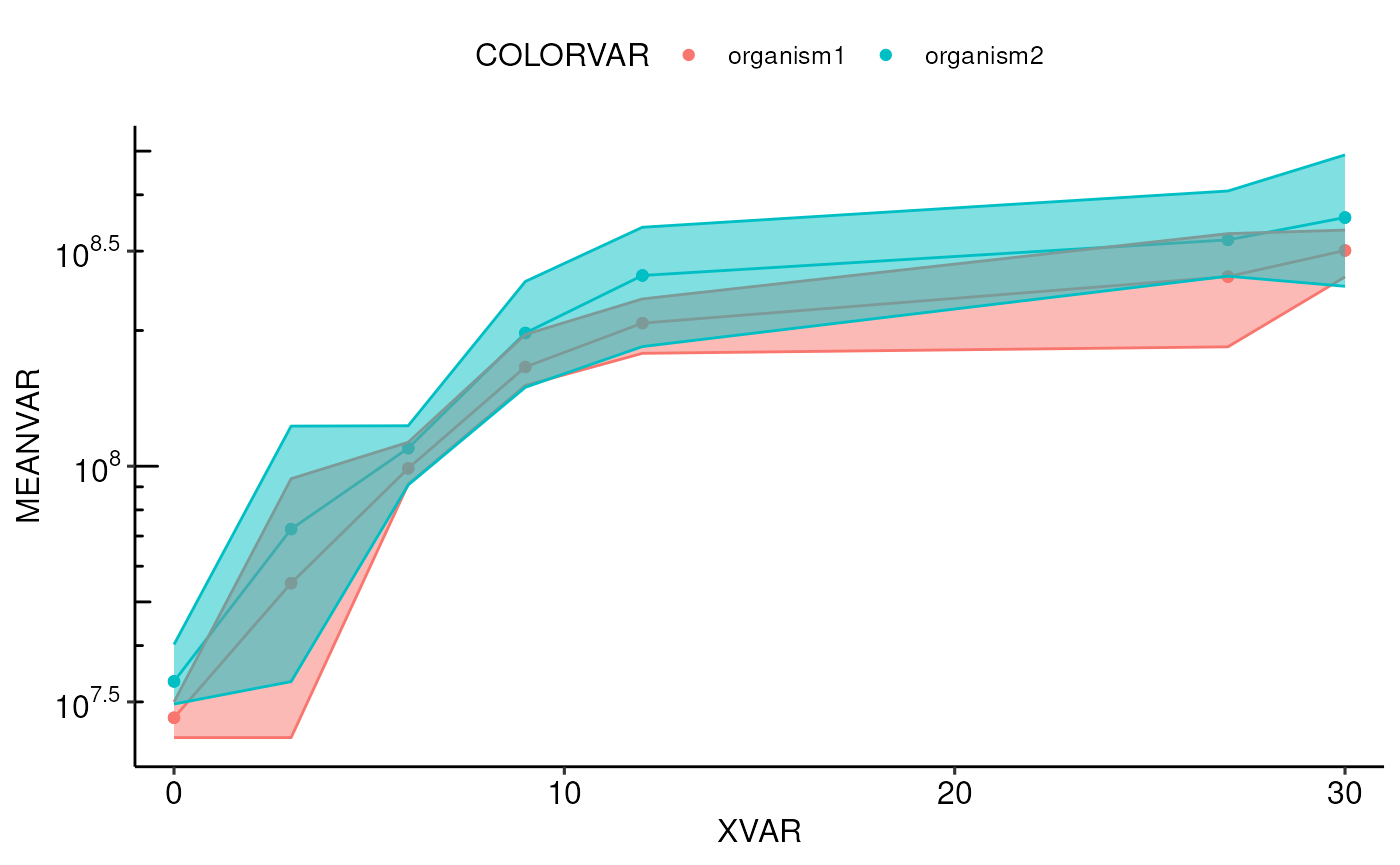
Modify your growth curve plot
Since the plotGrowthCurve function is basically just a
wrapper around a ggplot2 plot with some default settings, you can modify
the plot to your liking. Here is one example:
library(ggplot2)
archaeacentre::plotGrowthCurve(testData, timepoint, concentration, grouping = c("timepoint", "organism"), organism, type = "robert") +
ggplot2::labs(title = "Growth curve of some test data", x = "Timepoint after inocculation in [h]", y = "Concentration in [cells/mL]") +
ggplot2::guides(color = guide_legend(title = "Species")) +
ggplot2::facet_grid(~organism)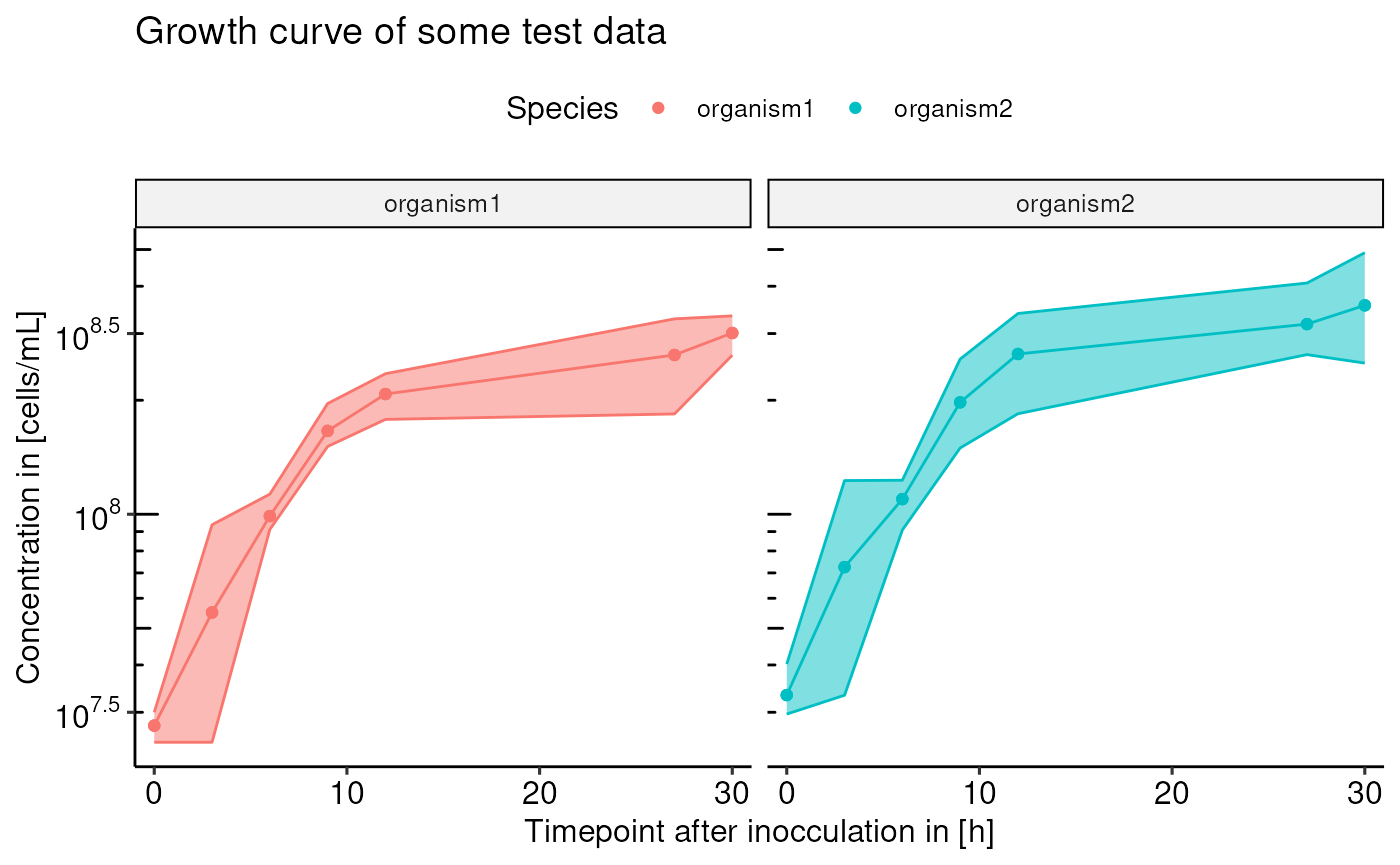
Get PDB Annotation data for a given PDB filename as returned by Foldseek
is a fast search tool for comparing protein structures. When searching for similar structures in the PDB, Foldseek returns a table which contains the “target” column. In the case of searches against the PDB, this target column contains the PDB filename of the hit, which is not easily interpretable.
The provides structural and functional annotations for macromolecules stored in the Protein Data Bank (PDB).
The get_pfam_annotation_for_targets() function automates
the extraction of Pfam domain annotations for these target PDB filenames
returned by Foldseek,using the .
# Get a vector of PDB filenames. This could be the "target" column of a Foldseek search result table.
targets <- c("1U04_assembly1.cif_A", "8HL4_assembly1.cif_L18P")
# Get the Pfam annotations for these targets. As the RCSB API has a rate limit, we recommend to use a batch size of 1000 or lower.
pfam_results <- get_pfam_annotation_for_targets(targets, batch_size = 1000)
head(pfam_results)
#> # A tibble: 2 × 4
#> target rcsb_id title pfam_description
#> <chr> <chr> <chr> <chr>
#> 1 1U04_assembly1.cif_A 1U04.A Crystal structure of full le… Argonaute, N-te…
#> 2 8HL4_assembly1.cif_L18P 8HL4.Q Cryo-EM Structures and Trans… Ribosomal large…